Yup, the docs fragmentation is a serious problem. Since there are so many entrypoints, I think we are going to have to be okay with some amount of information duplication, since people are going to arrive from so many different directions.
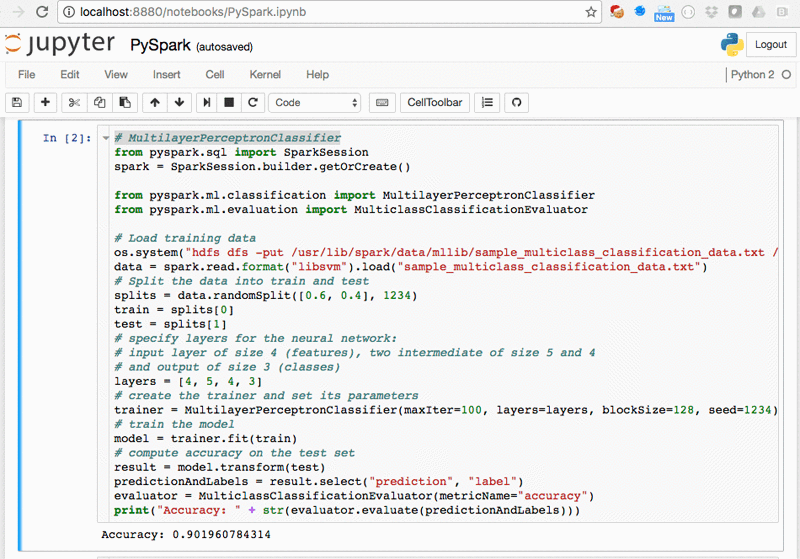
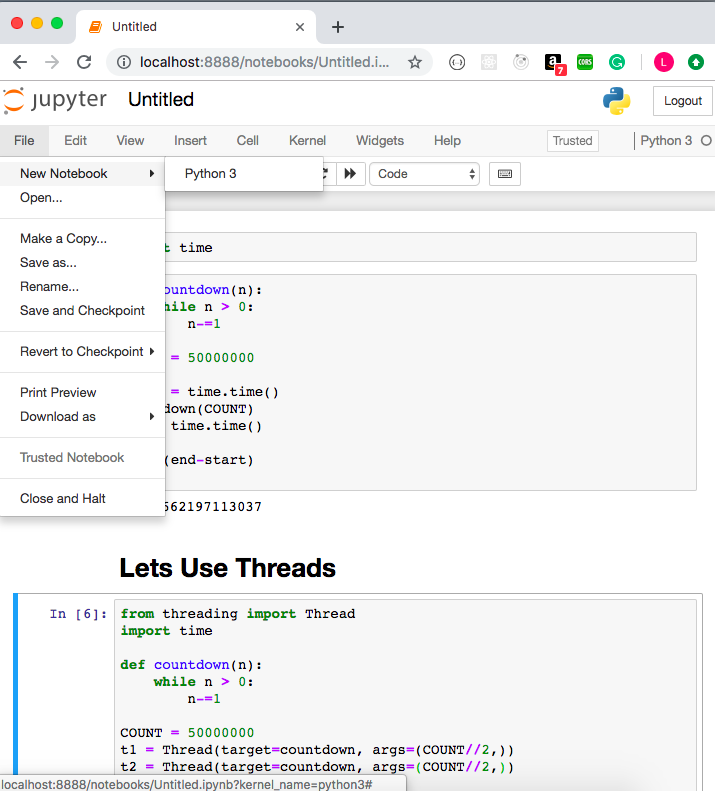
Try Jupyter with Python. A tutorial introducing basic features of Jupyter notebooks and the IPython kernel. Try JupyterLab. JupyterLab is the new interface for Jupyter notebooks and is ready for testing. Give it a try! Try Jupyter with Julia. A basic example of using Jupyter with Julia. Now when you login to jupyter webserver, you can see the Python 2 menu item in the New drop down list. Click this menu item will create a notebook that can submit Python 2 source code to jupyter web server and start a ipython kernel process to run the python 2 source code.
Assuming you are using pip, you can install the
ipykernel package with Python 2, and then install the kernelspec, so Jupyter knows how to start the kernel (no real difference if you use conda):Installation of packages such as numpy, etc. is unaffected by Jupyter. The only additional step is installing the kernelspec after installing the kernel package itself, so that Jupyter can find the kernel. JupyterHub itself also has no effect on installation, other than the fact that you presumably want to do system-wide installs of packages and kernelspecs (generally the default already) so that all Hub users have access to what you install.
A totally complete installation of JupyterHub, notebook servers, and kernels for both Python 2 and 3 (sudo may be required, depending on the system):
And you can install whatever packages you want for Python 3 and/or Python 2 in the usual ways.
If you do the same with Anaconda, it's a bit easier to get all the way to a full scientific Python stack with multiple versions of Python:
I'll add something like this to the JuptyerHub docs.
I have installed Jupiter notebook and I only have python 2 as a default kernel. I want to change it from python 2 to python 3. How could I do that?
Satyam KumarSatyam Kumar
3 Answers
Follow the link for managing python. If you use python 2, then install python 3 by using this command.
After installing python 3, activate python 3 by using this command
Then open jupyter notebook, you will find python on your kernel.
Sadman SobhanSadman Sobhan
You can do this with the following steps:
conda create -n py36 'python=3.6' ipykernel #Replace 3.6 with desired version.- To activate installed jupyter kernal you need run,
source activate py36 python -m ipykernel install --user- The interesting part: if you want to switch between kernels (py2-py3) in the same notebook, you need to run,
conda install nb_conda
However, if at any point you realize that some of the modules are not available, for that you need to check Anaconda Python version.
if it is not python 3.x, you need to run
I hope it helps and enjoy switching python versions in the same notebook. You can try to print something in both python2 and python 3.
Kripalu SarKripalu Sar
In case you installed the python/jupyter with anaconda, simply update the python from 2.* to 3.*
xmindataxmindata